Welcome to the Population Genomics Laboratory
Department of Genetics, Genomics and Informatics

We are interested in understanding causes and consequences of genetic diversity and how natural selection in humans affects loci related to diseases.
Fascinating (Lieutenant Spock)
Research
Population Genomics and Health Equity
The Biorepository and Integrative Genomics (BIG) Initiative in Tennessee
The Biorepository and Integrative Genomics (BIG) Initiative in Tennessee has developed a pioneering resource to address gaps in genomic research by linking genomic, phenotypic, and environmental data. BIG is recruiting participants mainly from Memphis, TN, with plans to include a total of 100,000 samples over the next five years.
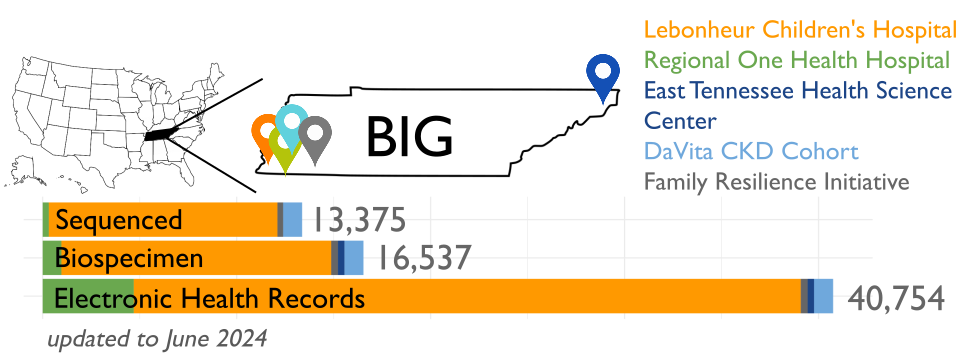
-
BIG is partnering with the Genomic Information Commons, a consortium of top children’s hospitals, to conduct genomics research aimed at discovering the genetic foundations of human disease in diverse populations.
-
BIG is part of the Togheter for Change initiative (T4C)
Characterizing Genetic Diversity in the BIG Cohort
Analysis of 13,152 genomes from BIG revealed significant genetic diversity, with 50% of participants inferred to have non-European or various types of admixed ancestry, highlighting the importance of inclusive genomics in understanding health equity.
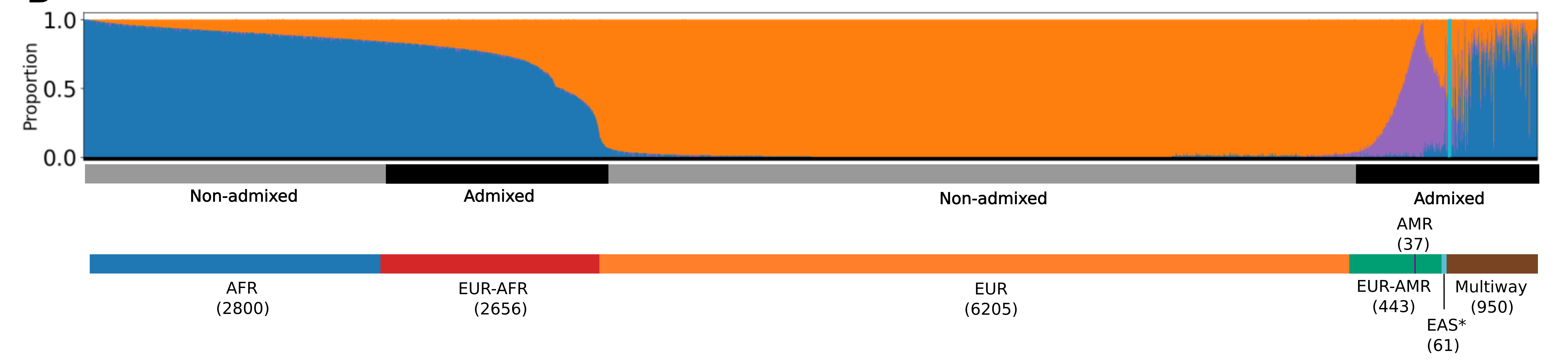
Global ancestry deconvolution of 13,152 sequenced individuals, based on reference populations in the 1000 Genomes and HGDP data sets. Each vertical bar represents one individual, colors are proportional to inferred ancestry. Individuals were further grouped based on the ancestry proportions in seven categories based on the likelihood of similarity to reference populations (AFR= Africans; EUR-AFR = Europeans Africans admixed; EUR = Europeans; EUR-AMR = Europeans Americans admixed; Multiway = more than two ancestries; number of individuals per category in parentheses). See the open access paper.
Identity-by-Descent as a Window into Shared Environment and Health
We use identity-by-descent (IBD) sharing patterns in the BIG cohort to identify recent shared ancestry and fine-scale population structure. These patterns help connect inherited genomic segments with shared environmental exposures and disease risk.
Population Pangenomics
A pangenome is a comprehensive collection of all the genetic variation present in a species, which overcomes the limitations of reference-based genomics by including both common and rare genetic variations in a single reference genome.
The Human Pangenome Reference Consortium aims to sequence 300 people to create a pangenome of 600 haplotypes and has currently released a first draft of the human pangenome reference based on 47 phased diploid assemblies from a group of genetically diverse individuals
Uncharted Genomes
Our team evaluated, for the first time, human population structure using markers from short arms of the acrocentric chromosomes [PMID: 37165242]. We found that markers from these regions have less power to distinguish populations compared to other regions. This is consistent with the understanding that short arms of acrocentric chromosomes undergo recombination between non-homologous chromosomes, similar to the X and Y pseudohomologous regions [PMID: 37165241]. Our findings on the patterns of linkage disequilibrium in these regions support this idea.
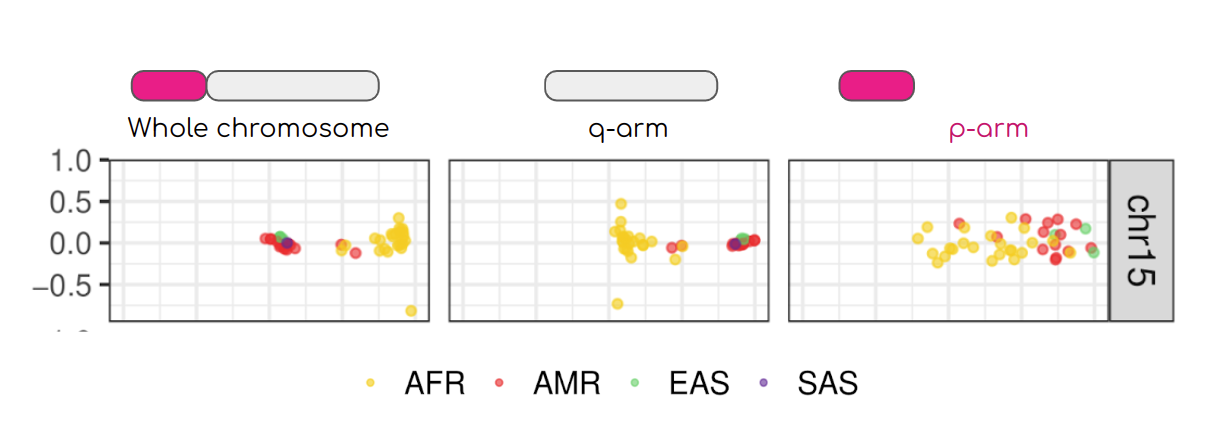
Well-known population stratification is not visible in the p-arms of acrocentric chromosomes. This observation is compatible with recombination occurring between the p-arms of heterologous acrocentric chromosomes. Here is an example of chromosome 15, PCA with markers from the whole chromosome, only the q-arm, only the p-arm. AFR: Africans; AMR: Native Americans; EAS: East-Asians; SAS: South-East Asians.
Pangenomics tools for population genetics
Bat’s pangenomics
Genomics of early embryonic development
How natural selection acts on early human development
We investigate adverse outcomes of embryonic development like recurrent pregnancy loss and preeclampsia to identify the genetic factors that influence reproductive outcomes and pregnancy complications. This knowledge furthers our understanding of human evolution and informs efforts to improve pregnancy outcomes.
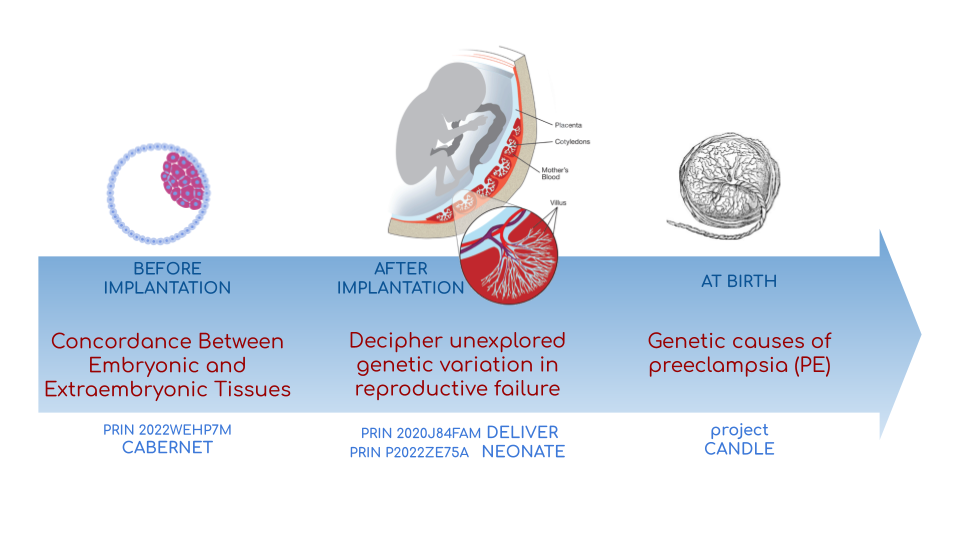
In CABERNET we aim to determine the extent of chromosomal mosaicism
between embryonic and extraembryonic tissues using single cell DNA
sequencing.
In DELIVER / NEONATE we want to identify genetic factors contributing
to reproductive failure and recurrent miscarriage. We will use single
cell strand sequencing to map balanced rearrangements and whole genome
sequencing of euploid miscarried embryos to identify causative
variants.
In CANDLE we want to uncover gene expression patterns associated with
APOL1 risk alleles and preeclampsia in African American women. We will
examine the role of ancestry in mediating the relationship between
APOL1 genotype and preeclampsia risk. The results can provide insights
into genetic and molecular basis of preeclampsia.
This project is in collaboration with Francesca Antonacci University of Bari Aldo Moro; Carlo Alviggi, University of Naples Federico II; Marcella Vacca, and Gabriella Lania, National Research Council
Funding
-
PRIN 2020J84FAM Ministero dell’Universita e della Ricerca
-
PRIN 2022WEHP7M Ministero dell’Universita e della Ricerca
-
PRIN P2022ZE75A Ministero dell’Universita e della Ricerca


Current Members
Vincenza Colonna

Associate Professor, Department of Genetics, Genomics and Informatics
website
University of Tennessee Health Science Center, Memphis, TN
Director, Integrative Genomics Biorpository, Department of
Pediatrics
Children’s Foundation Research Institute, Memphis, TN
Senior Researcher (on leave of absence), National Research Council
website
Institute of Genetics and Biophysics, Naples, Italy
I graduated in Evolutionary Biology from the University of Naples Federico II and did postdoctoral research at the University of Ferrara (Italy) and at Wellcome Trust Sanger Institute in Cambridge (UK). I was lectures in Genetics and Bioinformatics at the University of Ferrara (Italy). I am now leading the Population genomics laboratory at the University of Tennessee, College of Medicine, in the Department of Genetics, Genomics and Informatics.
I am a genomicist and an expert in human evolutionary and population genomics and bioinformatics. In my postdoctoral research I was part of the international consortium 1000 Genomes[PMID: 26432245; 23128226] where I led contributions to two specific aspects. First, I contributed to develop FunSeq [PMID: 24092746], a tool that integrates non-coding information from relevant biological databases for the functional characterization of non-coding variants. Second, I lead a genome-wide scan to identify genomic regions with exceptionally high levels of population differentiation [PMID: 24980144] demonstrating that these regions are enriched for positive selection events and that one half may be the result of classic selective sweeps. Findings from both sub-projects have since been applied to demographic inference and the molecular diagnosis of cancer and myeloid malignancies [PMID: 27121471, 22446628], and to deeper studies on positive selection at the ABCA12 gene [PMID: 30890716].
During my PhD I worked on human isolated populations contributing to characterize several isolated populations, describing the genomic consequences of isolation [PMID: 17476112, 19550436, 22713810], contributing to genetic association studies [PMID: 16611673, 18162505] and to characterize rare variation [PMID: 28643794]
I founded and led OBiLab, a project on training in Bioinformatics
Silvia Buonaiuto, Postdoctoral fellow

I am a postdoctoral researcher at the University of Tennessee Health Science Center in the Department of Genetics, Genomics, and Informatics. Prior to this, I worked as a postdoctoral researcher at the National Research Council in Italy. I earned a Ph.D. in Molecular Bioscience from the University of Campania "Luigi Vanvitelli." Before that, I completed my Master’s degree in Biology at the University of Naples Federico II, where I conducted my thesis in molecular biology at the Department of Biology.
I work on admixture mapping and pangenomics analysis in the BIG project, with a focus on connecting genotypes and phenotypes in admixed individuals through admixture mapping. I also lead sequence analysis for projects related to early embryonic development.
Franco Marsico, Postdoctoral fellow

I earned my degree in Biology from the University of Buenos Aires, Argentina, where I also completed my PhD in Computational Biology at the Calculus Institute. My research primarily focused on developing mathematical models for kinship inference, employing a Bayesian Approach. I am a postdoc in the Colonna lab, where my work centers on population genomics.
I am currently working on Admixture Mapping and Pangenomics Analysis in the Biorepository and Integrative Genomics (BIG) Initiative Cohort project. My focus is on studying recent natural selection signals in admixed populations. Additionally, I have a deep interest in evolution and how to compute processes that shape the history of life.
Laura Pignata, Postdoctoral fellow

I graduated in Molecular Biology at the University of Campania Luigi Vanvitelli, where I also completed my PhD program in Molecular Life Sciences. After the PhD I continued my work as researcher, focusing on the analysis of DNA methylation in imprinting disorders. I am visiting Dr. Colonna laboratory at the University of Tennessee, College of Medicine, in the Department of Genetics, Genomics and Informatics.
I analyze long-read sequencing data from human samples with mosaic methylation defects. My work focuses on identifying the mechanisms behind these defects by testing the roles of de novo mutations in cis and somatic recombination events that may arise during early embryogenesis.
Ernestine Amos-Abanyie, PhD Student

I have a master degree in Molecular Biology from the university of Ghana, I am a PhD candidate in the Biomedical Science program of the Department of Genetics Genomics and Informatics in co-supervision with Dr. Ashbrook.
I study mitochondrial DNA variation and its contribution to disease susceptibility in both the BXD mice and the BIG cohort.
Donna Vo, Research associate

I earned a Bachelor of Science in Microbiology at the University of Tennessee, Knoxville. I have a background in spoilage detection and mycology identification in the food and beverage industry.
I oversee sample collection and processing for the Integrative Genomics Biorepository that supports the BIG project.
Visiting
Giuseppe Comentale, PhD Student
Past Lab members
-
Flavia Villani, Master student, PhD student, 2019-2025
-
Gianluca Damaggio, Master student, PhD student, 2019-2023
-
Rosanna Maione, Research associate 2023
-
Madeleine Emms, Postdoctoral fellow, 2022-2023
-
Marialaura Zitiello, Master Student, 2022-2023
-
Antonella Mecca, Master Student, 2022-2023
-
Angela Sequino, Master Student, 2022-2023
-
Giuliana D’Angelo, Master Student, 2019-2020
-
Roberto Sirica, PhD student, 2015-2018
-
Gaia Leandra Cecere, undergraduate student, 2018
Past visiting
-
Dario Cannella, PhD student, visiting student, 2025
-
Angela Carfora, PhD student, visiting student, 2025
-
Davide D’angelo, Visiting master student, 2022
-
Marianna Buonaiuto, visiting Postdoc, 2017
-
Lucia De Martino, visiting master Student, 2016
Publications
See them on Google Scholar
Contacts
Vincenza Colonna, PhD enza.colonna@gmail.com
-
University of Tennessee Health Science Center, TSRB room 405
71 S Manassas St, Memphis TN 38163
map vcolonna@uthsc.edu website at UTHSC -
Children’s Foundation Research Institute
50 N. Dunlap St. Memphis, TN 38105
map -
Istituto di Genetica e Biofisica "Adriano Buzzati-Traverso" piano R, stanza 11
via Pietro Castellino 111 - 80131 Napoli - Italy
map vincenza.colonna@igb.cnr.it